Note
Click here to download the full example code
7.11. Path and skimming¶
On this example we show how to perform path computation and network skimming for the Sioux Falls example model.
## Imports
from uuid import uuid4
from tempfile import gettempdir
from os.path import join
from aequilibrae.utils.create_example import create_example
# We create the example project inside our temp folder
fldr = join(gettempdir(), uuid4().hex)
project = create_example(fldr)
import logging
import sys
# We the project open, we can tell the logger to direct all messages to the terminal as well
logger = project.logger
stdout_handler = logging.StreamHandler(sys.stdout)
formatter = logging.Formatter("%(asctime)s;%(levelname)s ; %(message)s")
stdout_handler.setFormatter(formatter)
logger.addHandler(stdout_handler)
imports
from aequilibrae.paths import PathResults
we build all graphs
project.network.build_graphs()
# We get warnings that several fields in the project are filled with NaNs. Which is true, but we won't use those fields
2023-01-16 13:35:10,038;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
2023-01-16 13:35:10,072;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
2023-01-16 13:35:10,106;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
2023-01-16 13:35:10,139;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
2023-01-16 13:35:10,172;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
2023-01-16 13:35:10,206;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
we grab the graph for cars
graph = project.network.graphs["c"]
# we also see what graphs are available
# project.network.graphs.keys()
# let's say we want to minimize distance
graph.set_graph("distance")
# And will skim time and distance while we are at it
graph.set_skimming(["free_flow_time", "distance"])
# And we will allow paths to be compute going through other centroids/centroid connectors
# required for the Sioux Falls network, as all nodes are centroids
# BE CAREFUL WITH THIS SETTING
graph.set_blocked_centroid_flows(False)
/home/runner/work/aequilibrae/aequilibrae/aequilibrae/paths/graph.py:445: FutureWarning: The default value of numeric_only in DataFrameGroupBy.sum is deprecated. In a future version, numeric_only will default to False. Either specify numeric_only or select only columns which should be valid for the function.
df = self.__graph_groupby.sum()[[cost_field]].reset_index()
/home/runner/work/aequilibrae/aequilibrae/aequilibrae/paths/graph.py:479: FutureWarning: The default value of numeric_only in DataFrameGroupBy.sum is deprecated. In a future version, numeric_only will default to False. Either specify numeric_only or select only columns which should be valid for the function.
df = self.__graph_groupby.sum()[skim_fields].reset_index()
instantiate a path results object and prepare it to work with the graph
res = PathResults()
res.prepare(graph)
# compute a path from node 8 to 13
res.compute_path(8, 4)
# We can get the sequence of nodes we traverse
res.path_nodes
array([8, 9, 5, 4])
# We can get the link sequence we traverse
res.path
array([21, 23, 11])
# We can get the mileposts for our sequence of nodes
res.milepost
array([ 0. , 2213.32449172, 3927.86012613, 5188.91313598])
# If we want to compute the path for a different destination and same origin, we can just do this
# It is way faster when you have large networks
res.update_trace(13)
res.path_nodes
array([ 8, 9, 10, 15, 22, 21, 24, 13])
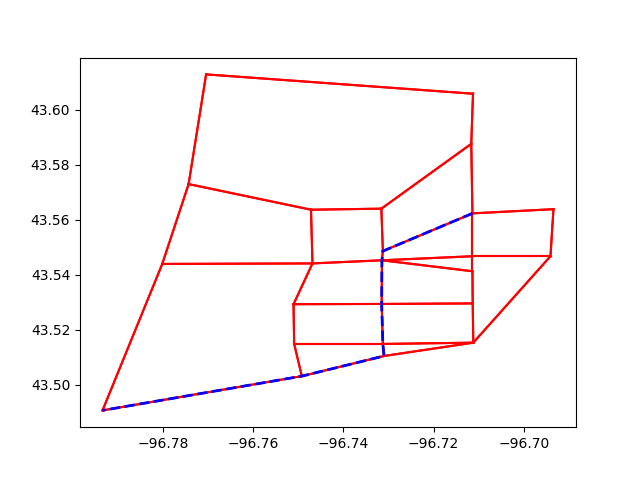
If you want to show the path in Python We do NOT recommend this, though…. It is very slow for real networks
import matplotlib.pyplot as plt
from shapely.ops import linemerge
links = project.network.links
# We plot the entire network
curr = project.conn.cursor()
curr.execute("Select link_id from links;")
for lid in curr.fetchall():
geo = links.get(lid[0]).geometry
plt.plot(*geo.xy, color="red")
path_geometry = linemerge(links.get(lid).geometry for lid in res.path)
plt.plot(*path_geometry.xy, color="blue", linestyle="dashed", linewidth=2)
plt.show()
## Now to skimming
from aequilibrae.paths import NetworkSkimming
But let’s say we only want a skim matrix for nodes 1, 3, 6 & 8
import numpy as np
graph.prepare_graph(np.array([1, 3, 6, 8]))
2023-01-16 13:35:10,355;WARNING ; Field(s) name, lanes has(ve) at least one NaN value. Check your computations
# And run the skimming
skm = NetworkSkimming(graph)
skm.execute()
# The result is an AequilibraEMatrix object
skims = skm.results.skims
# Which we can manipulate directly from its temp file, if we wish
skims.matrices
array([[[0.00000000e+00, 0.00000000e+00],
[6.00000000e+00, 4.84008627e+03],
[9.00000000e+00, 6.46536178e+03],
[1.30000000e+01, 8.88889720e+03]],
[[6.00000000e+00, 4.84008627e+03],
[0.00000000e+00, 0.00000000e+00],
[9.00000000e+00, 6.73286110e+03],
[2.10000000e+01, 6.82126560e+03]],
[[9.00000000e+00, 4.97310839e+03],
[1.00000000e+01, 8.30423993e+03],
[0.00000000e+00, 0.00000000e+00],
[4.00000000e+00, 2.42353542e+03]],
[[1.10000000e+01, 6.23416140e+03],
[2.00000000e+01, 8.12432422e+03],
[2.00000000e+00, 1.26105301e+03],
[0.00000000e+00, 0.00000000e+00]]])
# Or access each matrix
skims.free_flow_time
array([[ 0., 6., 9., 13.],
[ 6., 0., 9., 21.],
[ 9., 10., 0., 4.],
[11., 20., 2., 0.]])
# We can save it to the project if we want
skm.save_to_project("base_skims")
# We can also retrieve this skim record to write something to its description
matrices = project.matrices
mat_record = matrices.get_record("base_skims")
mat_record.description = "minimized FF travel time while also skimming distance for just a few nodes"
mat_record.save()
2023-01-16 13:35:10,407;WARNING ; Matrix Record has been saved to the database
project.close()
Total running time of the script: ( 0 minutes 0.519 seconds)